Science as a vocation
Max WEBER
Information on the influenza virus
This information was provided "as is" during the H7N9 avian flu outbreak in China in years following the 2013 epidemic
Starting on February 19, 2013, two patients with severe pneumonia and pulmonary distress were admitted to Shanghai Fifth Hospital affiliated with Fudan University. The first patient, an 87-year-old man had become ill on February 19th and died on February 27th. The second patient, also male and 27 years old, became ill on February 27th and died on March 4th. A study carried out at the carried out at the Laboratory of Pathogen Diagnosis and Biosafety of Shanghai Public Health Clinical Center, identified that the disease was caused by a previously unknown influenza virus coming from reassortment between three parent viruses. Remarkably, the patients had only been in contact with pigs. Several similar diseases with life-threatening symptoms followed, some of them having been in contact with poultry.
Infection by an influenza A virus of strain H7N9 was confirmed on march 10th. On march 28th 2013 a 74-old man became ill and his condition worsened rapidly. On april 7th, 21 persons had been identified as infected by the virus- ten in Shanghai, six in Jiangsu, three in Zhejiang and two in Anhui. Four patients in Shanghai and two in Zhejiang have died. On april 10th 33 persons have been recognized to be infected by the virus and nine died of the disease. At the end of april the epidemic appeared to level off. This can be expected with the coming of spring, as well as the closure of live poultry markets. However the exact origin of the disease is not well established. The virus appears to derive from a reassortment of four different avian flu viruses: H9N2, H7N3, H4N9, H11N9.
As the epidemic menaces to reappear in autumn it may be interesting to remember the situation with the better known epidemic of avian flu caused by the H5N1 virus. A short epidemic, back in 1997 infected 18 Hong Kong residents in 1997, killing 6. A similar but not identical virus reappeared in 2001.
Since then it has been lingering in many countries, being endemic in Indonesia, in China, in other East Asia countries, as well as in Egypt. It remains a matter of great concern because of its apparent letality, but it has not yet generated consistent person to person contamination. Recently several laboratory studies have caused a considerable controversy when experiments were developed in the laboratory to see whether it was possible to propagate a self-sustained form of the virus that would infect ferrets, animals that are used as reliable models of the human disease. The outcome of the experiment was positive, demonstrating that, indeed, we may expect the virus to become infectious via mammalian contacts, between humans in particular. In this context it is noteworthy that an isolate of the H7N9 virus has already been found to be able to infect ferrets.
Flu is an old disease of ducks (usually mild) and the usual transmission route is from duck to pig to man. In the case of the H7N9 (2013) virus the disease does not cause symptoms, or only mild symptoms in birds. Why such a bird disease should infect humans? Typically, most of our diseases, while ultimately due to a pathogen, result from our social behaviour. The influenza virus is often coming from China. This is likely because of the structure of Chinese farms (remember that the Chinese character for family is the pig under a roof 家)... The disease becomes dangerous when the transmission shifts directly from man to man.
A severe outbreak appears every couple of decades or so ("Asian flu", "Hong Kong flu" etc...), that progressively invades the whole world, while the virus mutates to adapt to the reaction of the human immune system on a more or less yearly basis. After somebody has got the disease he or she usually recovers, with a strong immune response. But this selects mutant viruses with the following trick: the antibodies of the host adapt closely to the shape of a protein covering the virus; but one way for the virus to avoid being recognized is to change one motif of that protein to a bigger motif. In this situation the antibodies are more or less inefficient. During this adaptation time people are however somewhat protected by the past infections and the disease is not so dangerous. After some time, the pool of variation is completely covered, and the only way out for the virus is to undergo more drastic changes, such as reassortment with similar viruses. Thus, in general, one finds a moderately severe disease every quarter of a century, followed by "ordinary" outbreaks every year as the rule.
Now, there are sometimes new trends, corresponding to the spreading of a form that is severe in birds (bird flu) rather than mild. This severe disease can directly infect man: this is the case of H5N1, and recently of H7N7. If that disease can adapt to transmit from man to man, then this will create an extremely dangerous outbreak, because no preexisting immunity is present in the human population. This is why the WHO is concerned at present, because this event will most probably happen in a not so remote future. The reason is the following: the flu virus is made of a collection of independent fragments. Each one can mutate, producing a form that may escape the immune system of the host. But the most dangerous event happens when two different flu viruses infect a given animal (man included). Indeed, in this situation the fragments may reassort and produce all kinds of viruses, some adapted to multiplication in the human host, with a disguise allowing it to escape its immune system. It is most likely that this will happen sooner or later. The only way out is to try to guess what will happen, and prepare a vaccine (ie preset the immune system by vaccination) against the new virus... This is today's challenge.
Influenza A viruses (family Orthomyxoviridae) have a genome of eight negative sense single-stranded RNA segments that encodes up to 13 proteins.
The influenza virus (or virion, in the medical jargon) is spherical in shape, with a rough surface. It is an enveloped virus - this means that the outer layer is a membrane made of fatty compounds (lipids) that are taken from the host cell in which the virus multiplies. It is covered by a coat of "sugared" proteins (that is, proteins tagged with particular sugars, named glycoproteins), with appendages ("spikes") that protrude out of the virus and allow it to bind on specific receptors of the surface of some of the host cells. These proteins are named HA (hemagglutinin) and NA (neuraminidase). Hemagglutinin serves as an attachment to the recipient cell, that triggers internalization of the virus. Neuraminidase, by constrast is mainly used by the virus, while formed in the cell's interior (cytoplasm), to exit the cell, triggering a new cycle of infection. It is an enzyme that cleaves sialic acid residues (acidic sugars tagging proteins) from internal receptors for the virus, enabling the virus to spread throughout the body.
These proteins determine the subtype of influenza virus (A/H5N1, for example). There are 16 subtypes of HA protein and 9 subtypes of NA proteins detected in wild birds. More subtypes exist if one considers also the type B influenza viruses as well as possible other reservoirs. Because they are located at the surface of the virus the HA and NA are important in the immune response against the virus; antibodies (proteins made by us to combat infection) against these spikes may protect against infection. Being an enzyme, neuraminidase can be inactivated by antiviral drugs. Relenza and Tamiflu are well-known example of such neuraminidase inhibitors.
Beside proteins making the spikes, others are embedded in the lipid membrane, and some make a scaffold that support the overall shape of the virion. The M2 protein lies within the membrane, with some of it exposed at the surface. It is an ion channel important for regulating the osmotic pressure inside the virion. It is the target of the antiviral compounds such as the once used adamantanes, which clog the channel or enter the virion where they may interfere with the virus replicase. Among thoses, amantadine and rimantadine are no longer recommended as they appear to be inactive on most influenza virus strains. Beneath the lipid membrane is a viral protein called M1, or matrix protein. This protein, which forms a scaffolding shell, gives strength and rigidity to the lipid envelope. Within the interior of the virion are the viral RNAs. They make the genetic material of the virus; each of them code for one or two proteins. The interior of the virion also contains another protein, NS2, also named NEP, which is essential to orchestrate the export of the viral RNAs from the cell's nucleus, where they are synthesized. Overall the virus codes for 11 proteins.
Figure from doi:10.1371/journal.ppat.1000566
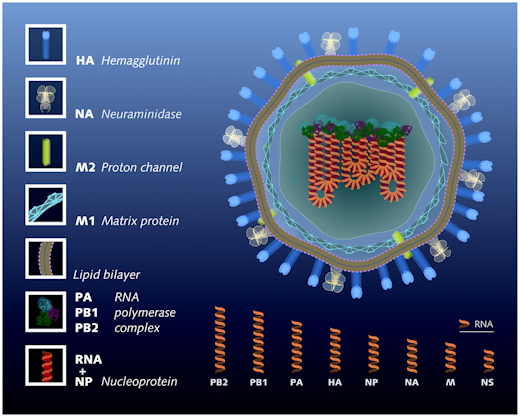
The longest codes for protein PB2, a component of
the viral gene that triggers initiation of RNA transcription and
replication
The next one codes for PB1, the elongating replicase/transcriptase
Next is PA, a protease associated to replicase/transcriptase
Then, HA, the hemagglutinin
Fragment NP codes for an RNA binding protein involved in RNA transport
Fragment NA, coding for neuraminidase, is essential for export of the
virus and release from the host cell
Then, the fairly short fragment M codes for proteins M1 and M2
Finally, the shortest fragment NS codes for protein NS1, involved in
RNA splicing and in translation, and protein NS2
These RNA segments are the genes of influenza virus. They are not used as direct templates for translation into proteins, but, by contrast, the are templates for messenger RNAs that are translated. This recognizes influenza A virus as an enveloped negative-strand RNA virus. The noncoding regions at the 3′ and 5′ termini of the eight vRNA segments contain highly conserved sequences that consist of 12 and 13 nucleotides, respectively. These sequences are partially complementary to allow the formation of a corkscrew structure that constitutes the viral promoter for replication and transcription.
There are now 16 subypes of hemagglutinin for type A influenza. Gulls are the primary hosts of H13 and H16 type viruses. The molecular basis for this host restriction is only partially understood. While the type refers to structural properties of the hemagglutinin, it may be associated systematically to variants of the other segments of the virus which might be responsible for host specificity.
Adamantanes were once used against influenza. Unfortunately uncontrolled used in veterinary medicine (aa often the case with other medications distributed by rogue veterinarians world-wide) probably resulted in selection of resistant virus strains. The consequence is that this class of drugs is now worthless.
The flu virus H7N9 is not totally unknown. It has been identified repeatedly all over the world, mostly infecting sea birds and ducks. Antiviral neuraminidase inhibitors might provide the first line of defense against a new flu virus. Drugs are therefore usually targeting the neuraminidase coded by the virus. It is expected that inhibition of this enzyme will stop the spread of the virus in the body and lead the infection to a dead end.Two inhibitors specific for neuraminidase are currently being used to control influenza infections. Zanamivir (4-guanidino-Neu5Ac2en) was invented in Australia and is marketed by Glaxo-SmithKline (Relenza). The other, Oseltamivir [4-acetamido- 5-amino-3-(1-ethylpropoxyl)-1-cyclohexene-1-carboxylic acid ethyl ester], was invented by Gilead Sciences and is marketed by Genentech (Hoffmann-LaRoche). Their target is the standard flu virus (H3N2), but explicit data on the targets and effects in vivo are not fully available.
Back in 2006 a study developed at the Department of Biological Sciences, BioCryst Pharmaceuticals, Inc., 2190 Parkway Lake Drive, Birmingham, AL 35244, USA showed that the drug Peramivir potently inhibited the neuraminidase enzyme N9 from the H1N9 virus in vitro. This is why this drug has been used early on, with the authorization of the highest authorities in China, despite the fact that it is still under phase III study.
Unfortunately the influenza virus is evolving very rapidly and it is likely that resistance will soon appear.
As in many diseases there may be a treatment using antibodies produced by recovered patients.
On may 1st, 2017, 1393 cases of human infection by the influenza virus H7N9 have been officially identified in China. The last patient died from the disease in Shanxi province on may 1st. Virulence of this influenza type was seldom known before march 2013, when several cases developed after infection of poultry in mainland China. Cases were essentially sporadic, with no identified clusters except in one recent case, in Tibet and a few earlier clusters. Fortunately, even in these cases, there was no person to person contamination. For several years the epidemic remained more or less stationary. However, since autumn 2016 the number of human cases seems on the increase (with respect to avian contamination), suggesting that the virus mutated to a more human-adapted form.
| Return to the Journalists page.... |